Native Mass Spectrometry (MS) Overview
Learn about native MS and the benefits and challenges of the emerging technology
What is native mass spectrometry (MS)?
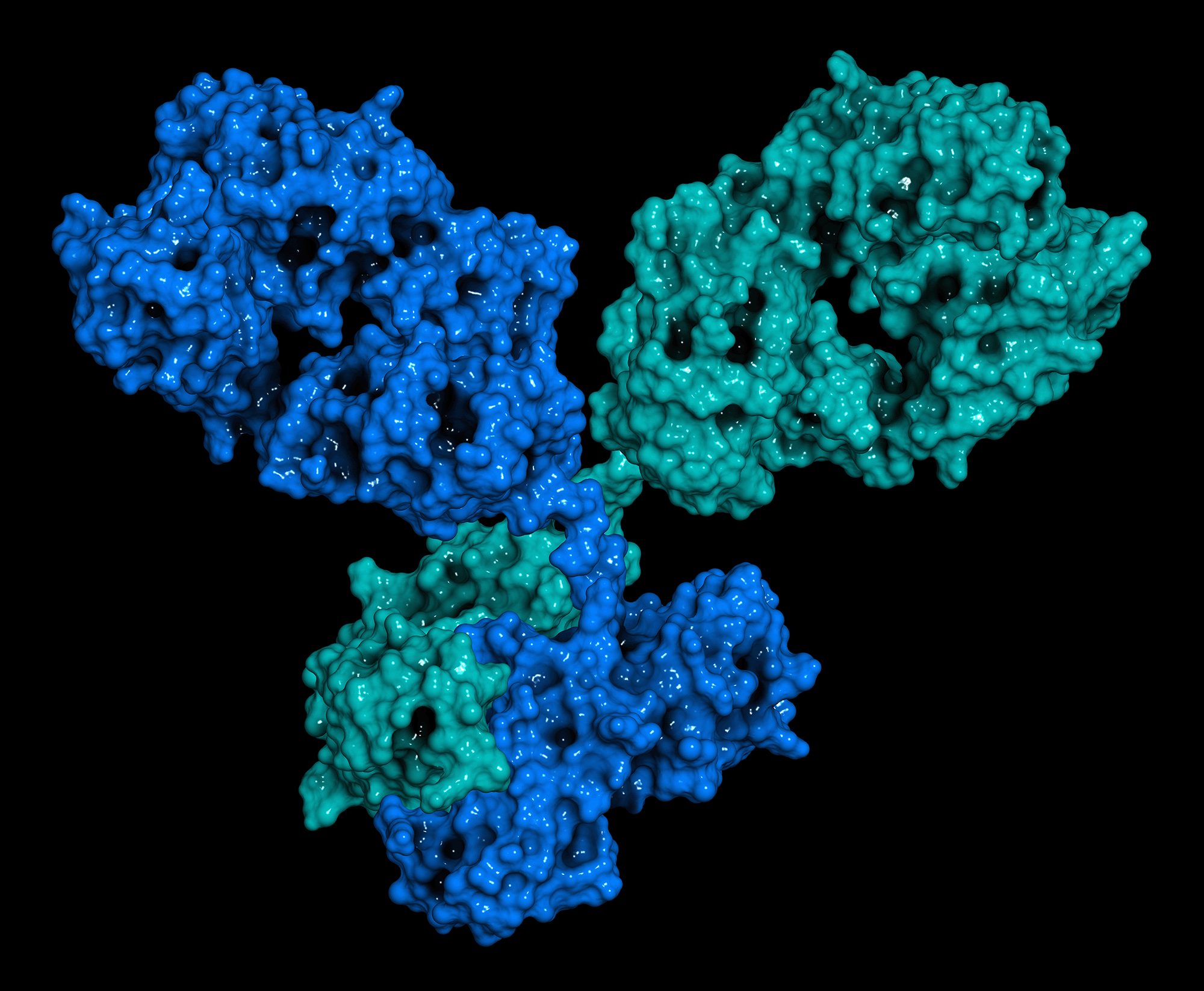
Mass Spectrometry (MS) is an analytical technique that is used to measure the mass to charge ratio of ions. Native MS (nMS) is an emerging approach for studying large biomolecules and their complexes in the solution via the electrospray ionization mass spectrometry (ESI-MS), while maintaining as much as possible the native structural features of the analytes and their interactions in the gas phase during the electrospray processes. The approach has gained significant traction in studying monoclonal antibodies (mAb), antibody-drug conjugates (ADCs), protein-protein complexes, protein-ligand interactions, and adeno-associated virus (AAVs), to name just a few.
Why is native MS important?
Native MS can disclose the composition, stoichiometry, kinetics, stability, and structure of subunits of proteins and protein complexes in a high-throughput manner. Improper protein folding can promote nonspecific protein aggregation by preventing their cofactor from attaching to them. As a result, many diseases can occur; that’s why it’s vital to examine these proteins’ structural components.
Benefits of native MS
- The fundamental benefit of Native MS is that it preserves the biomolecule’s native folded state, or in other words, the structure, and functionalities of the biomolecule prior to and during the ionization event. For example, nMS is used to study protein-protein complexes and protein-ligand binding for high-throughput drug screening, before in-depth functional and structural validation of the potential interactions.
- Native MS is also used for studying homogeneity, conformation (folded versus unfolded), and oligomerization state of biomolecules. All of these are important in basic biology research and new drug development.
- Finally, nMS does not demand a large sample in comparison to other conventional non-MS based techniques such as X-ray and CryoEM for structural biology methods; just a minimum of 10 picomoles is needed. Normally, for sample optimization and to run multiple analyses, 100-500 picomoles are used.
What is NS-MS?
The term ”NS-MS” has been proposed by our Founder & CEO, Daojing Wang, as an updated term for Native MS. He believes this better defines the current technological state of Native MS research and will lead to its mainstream adoption.
His argument was published in a September 2023 article for The Analytical Scientist.
From the article: “In my opinion, a better term for native MS might be “native spray mass spectrometry” (NSMS or NS-MS). This will remove the ambiguity surrounding what “native” means for the technique. Clearly defining which step is native – and decoupling sample preparation, the electrospray process, and mass spectrometry – will promote NSMS technology developments and adoption by a wider group of both MS and non-MS communities.”
Major fields impacted by native MS
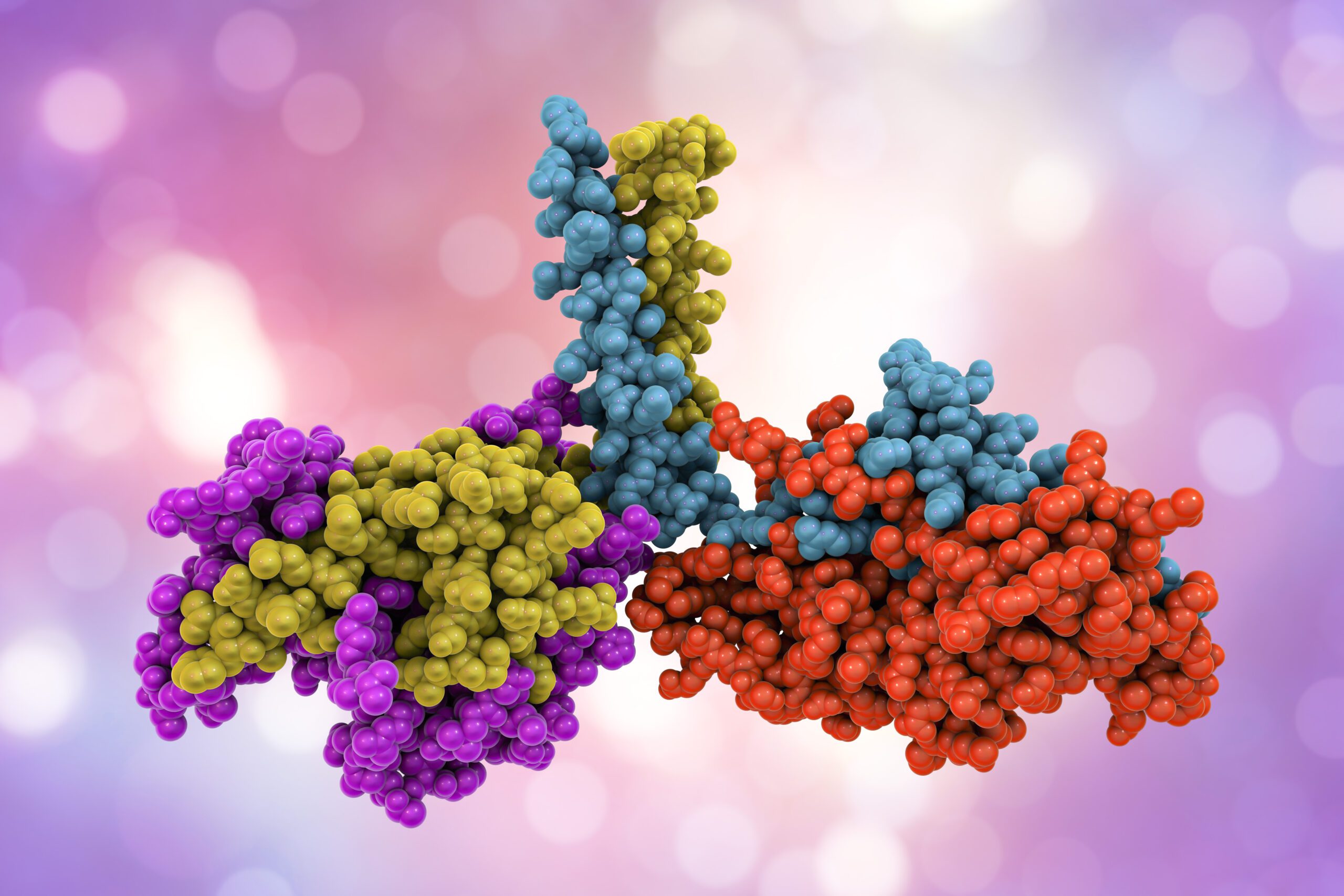
Structural biology research
Native MS can be used to understand the structure of macromolecules such as proteins, protein complexes, protein-ligand complexes, oligonucleotides, and post-translational modifications. nMS gives supplementary information to the existing structural biology methods, which do not provide information on the specific composition, structure, or dynamics of protein complexes. It is possible to analyze secondary, tertiary, and even quaternary structures of proteins and other biomolecules. Finally, protein-ligand interactions, lipid, and carbohydrate compounds, among other things, may be studied quantitatively and qualitatively.

Drug discovery and development
Proteins and enzymes are essential components of life processes and are linked to a variety of diseases, including cancer, diabetes, and Alzheimer’s disease. As a result, it’s critical to research and develop new pharmacological compounds that target specific proteins and protein-enzyme interactions.
Non-covalent interactions (ligand-target interactions) between small drug molecules and disease-related proteins regulate a variety of pharmacological processes in the treatment of various diseases. The development of analytical tools to evaluate such interactions, as native MS, can facilitate precision therapy of targeted illnesses, and targeted drug discovery.
Quality control of protein drugs
During the production process, protein drugs could be unfolded or modified. Therefore, it is important to monitor and characterize the purify and integrity of these drugs for downstream safety and efficacy studies. Native MS has been used to characterize the quality of protein drugs including the high molecular weight (HMW) size variants present in therapeutic monoclonal antibody (mAb) samples.
Why others fail to achieve native MS
Static nanospray ESI-MS
Typically, the samples are preloaded into a nanospray tip. This method is labor intensive for sample preparation and the tip is prompt to clogging. Furthermore, because there is no front end-separation by LC, nonspecific binding and/or impurity could be also detected.
High-flow ESI-MS
Typically, high temperature is required for high-flow ESI process, and this may break the native complexes or (partially) denature the proteins. Furthermore, high-flow ESI has a very low ionization efficiency, this dramatically reduces mass spectrometry detection sensitivity.
